To acknowledge the use of the Enlighten2 plugin and/or protocols, please cite the following publication:
Enlighten2: molecular dynamics simulations of protein-ligand systems made accessible
Aimed at:
- Experimental biochemists/enzymologists interested in gaining detailed insight into protein-ligand / enzyme-substrate complexes.
- Biomolecular researchers that would like to perform simulations in a high(er)-throughput fashion, e.g. for testing and hypothesis generation
Enlighten2 was developed with support from the UK Catalysis Hub [EPSRC grant EP/M013219/1] and BBSRC [BB/L018756/1 and BB/M026280/1].
Minimal software requirements
Package overview
Enlighten2 allows to easily prepare and run molecular dynamics simulations of protein-ligand systems. It requires no installation or knowledge of computational chemistry software and can be run on any machine (Windows, Mac or Linux) that has PyMOL and Docker installed. It consists of three main parts:
- The Python package containing a set of wrappers for AmberTools19 and Propka3.1.
- Docker image containing preinstalled Enlighten2 Python package, AmberTools19 and Propka3.1.
- The PyMOL plugin that provides a user friendly graphical interface to the Enlighten tools.
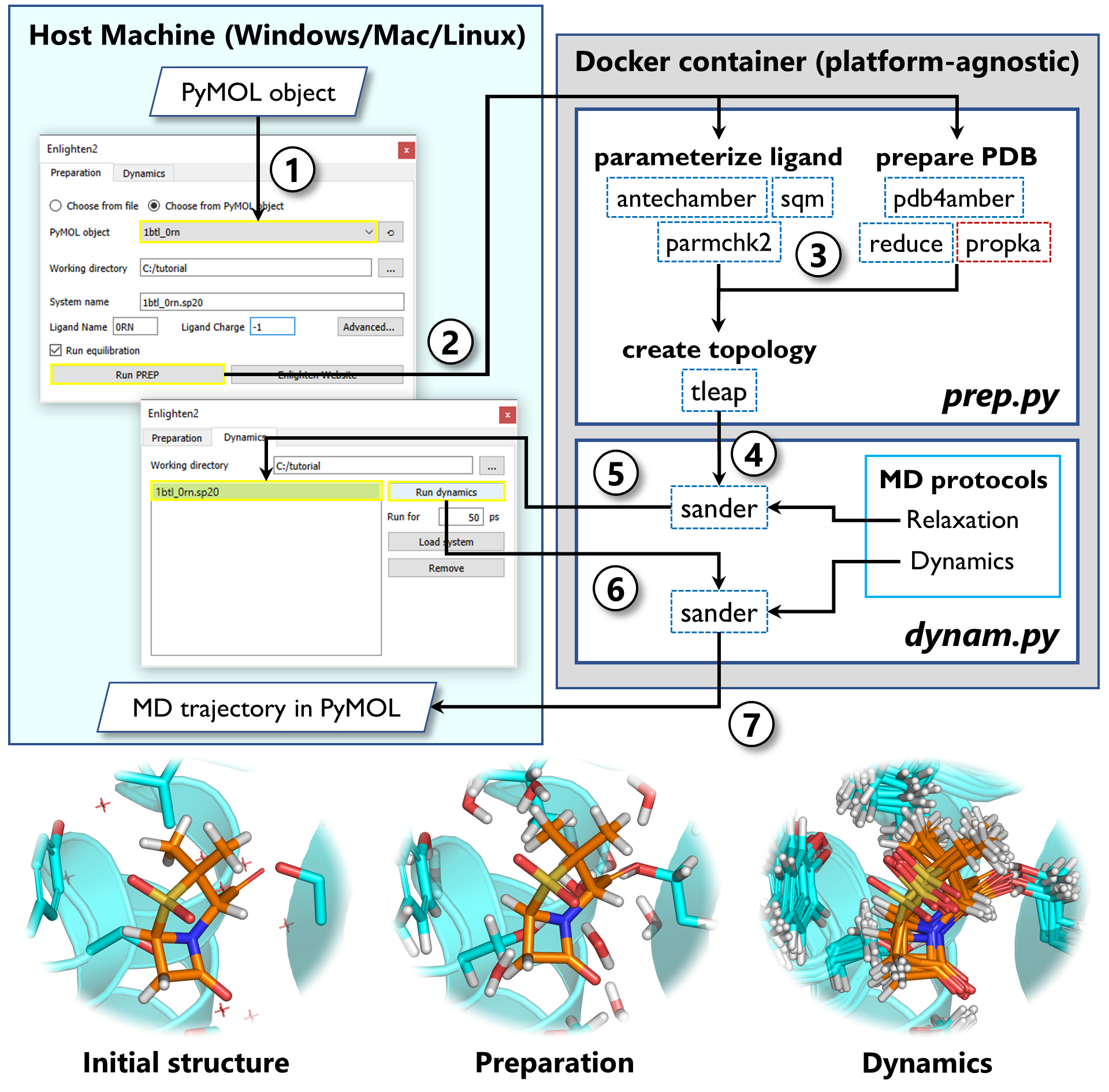
The Enlighten2 workflow
- Choose the PyMOL object in the Enlighten2 plugin and specify the system parameters (ligand charge, solvent sphere size etc.)
- Run preparation protocol. The plugin prepares all the input files and calls Docker to create the container from Enlighten2 image. If needed, the image is fetched automatically from DockerHub.
- Preparation protocol (prep.py) parameterizes the ligand, cleans up the protein structure and generated the simulation system.
- The prepared system is passed to molecular dynamics wrapper (dynam.py) to run the relaxation protocol.
- The relaxed system is passed back to PyMOL.
- Inspect the system and if there are no problems run the dynamics protocol for the desired simulation time.
- When the simulation is finished, the molecular dynamics trajectory is passed back to PyMOL.
Bugs in the package or plugin can be reported as an “Issue” in the corresponding GitHub repository (Python package, PyMOL plugin).
If you have feature requests, in-depth feedback or other thoughts about Enlighten that you would like to share, please get in touch.
24.06.2020: Fixed bug in the Docker image. If you tried to run the PyMOL plugin before this date, please run
docker image rm kzinovjev/enlighten2
in the Linux/MacOS terminal or Windows PowerShell.